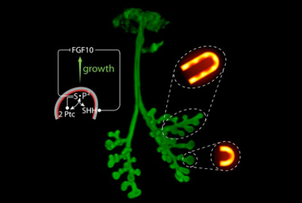
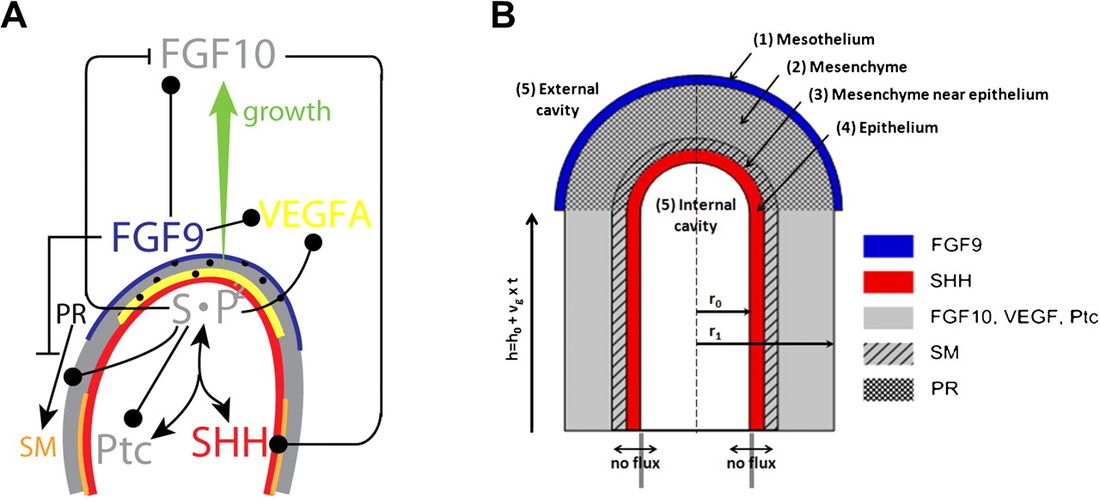
Many organs of higher organisms, such as the lung, kidney, pancreas, liver and glands are heavily branched structures. While many regulatory components and local interactions have been defined an integrated understanding of the regulatory networks that control the different branching processes is lacking. We propose that a Turing mechanism based on receptor-ligand interactions serves as a general mechanism for branching morphogenesis [1,2]. Unlike previously proposed Turing mechanism, ligand-receptor based Turing mechanisms permit pattern formation for a very large range of parameter values, which makes their evolution and re-use very likely [2,3].
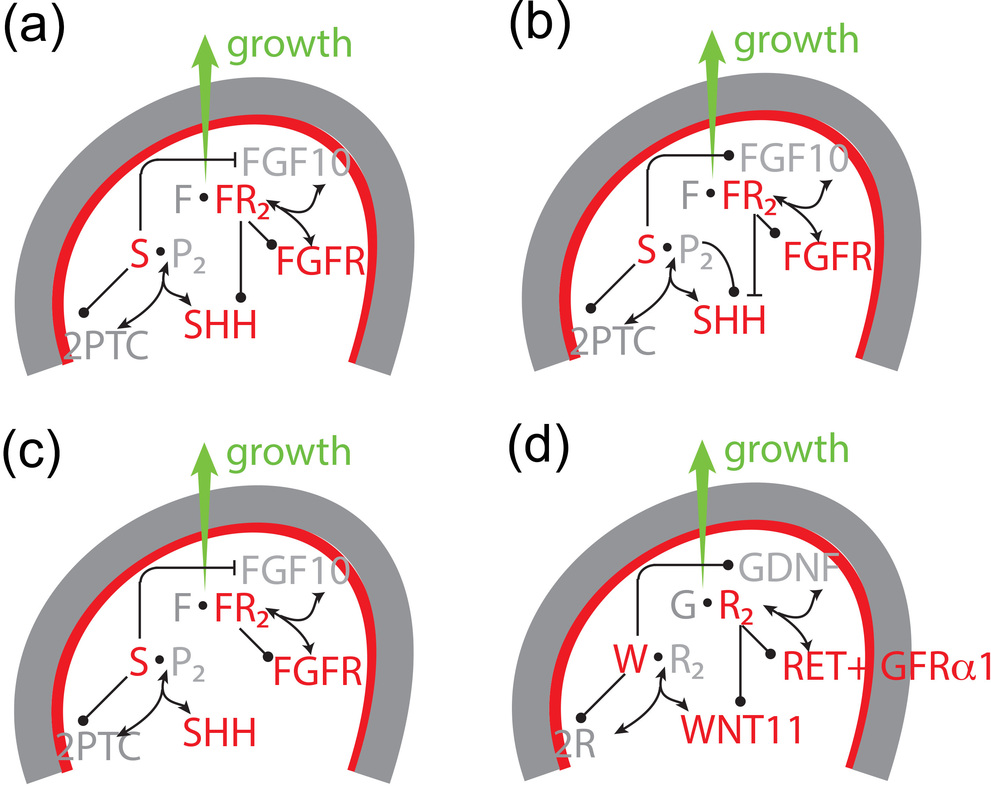
A general mechanism for branching morphogenesis?A similar regulatory network as in the lung (a) governs branching morphogenesis also in the prostate (a), salivary gland (b), and pancreas (c), and the identified mechanism is therefore likely to also apply to the development of these organs [1].
We further showed that also the GDNF-RET interaction (d), which controls branching in the kidney, gives rise to Turing patterns that reproduce the branching patterns both in wildtype and mutant kidneys [6]. We therefore propose that a Turing mechanism based on receptor-ligand interactions serves as a general mechanism for branching morphogenesis [1]. |
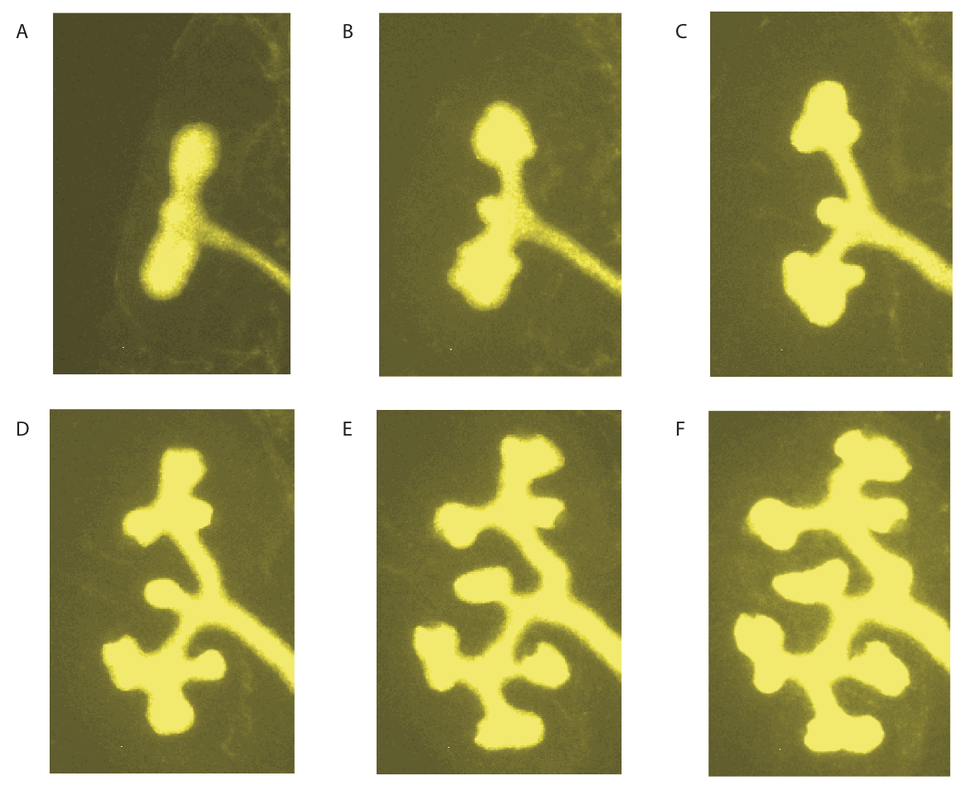
Current WorkTo further test our models for branching morphogenesis, we now solve these on realistic embryonic 3D domains that we obtain with optical projection tomography [2,7] as well as on movies that we obtain from organ culture [8,9]. We solve our models with the commercial FEM solver COMSOL [10-13].
|
PUBLICATIONS
- Iber D and Menshykau D, The Control of Branching Morphogenesis, Open Biology (2013) 3:130088
- Menshykau D, Blanc P, Ünal E, Sapin V and Iber D, An Interplay of Geometry and Signaling enables Robust Lung Branching Morphogenesis, Development (2014) accepted
- Kurics T*, Menshykau D* and Iber D (* shared first authors), Feedbacks, receptor clustering, and receptor restriction to single cells yield large Turing spaces for ligand-receptor based Turing models. Physical Review E (2014) 90, 022716
- Menshykau D, Kraemer C and Iber D Branch Mode Selection during early Lung Development. Plos Comp Biol (2012), 8, e1002377
- Celliere G, Menshykau D and Iber D A simple network is sufficient to control branch point selection, smooth muscle and vasculature formation during lung branching morphogenesis, Biology Open (2012), BIO20121339
- Menshykau D and Iber D Kidney Branching Morphogenesis under the control of a ligand-receptor based Turing mechanism. Phys. Biol. (2013) 10, 046003. The paper was selected as one of the Physical Biology's Highlights of 2013.
- Iber D, Tanaka S, Fried P, Germann G and Menshykau D Simulating Tissue Morphogenesis and Signaling, Springer Book Series: Methods in Molecular Biology. Book Title: Tissue Morphogenesis: Methods and Protocols. Editor: Celeste Nelson
- Adivarahan S, Menshykau D, Michos O and Iber D (2013) Dynamic Image-Based Modelling of Kidney Branching Morphogenesis. Lecture Notes in Computer Science (Springer), Computational Methods in Systems Biology (CMSB) 2013 IST Austria
- Schwaninger CA, Menshykau D, Iber D, Simulating Organogenesis: Algorithms for the Image-based Determination of Growth Fields. ACM Transactions on Modeling and Computer Simulation (TOMACS) (2014)
- Germann P, Tanaka S, Menshykau D, and Iber D (2011) Simulating Organogenesis in Comsol, Proceedings of COMSOL Conference 2011
- Menshykau D and Iber D, Simulating Organogenesis in Comsol: Deforming and Interacting Domains, Proceedings of COMSOL Conference, 2012
- Menshykau D, Germann P, Lemereux L and Iber D (2013) Simulating Organogenesis in COMSOL: Parameter Optimization for PDE-based models. Proceedings of COMSOL Conference 2013, Rotterdam
- Karimaddini Z, Ünal E, Menshykau D and Iber D, Simulating Organogenesis in COMSOL: Image-based Modeling, Proceedings of COMSOL Conference 2014, Cambridge